Hidradenitis suppurativa (HS) is a chronic inflammatory skin disease with poorly understood etiology. In a landmark genome-wide association meta-analysis (GWAS) involving 4,814 HS cases and over 1.2 million controls across five countries (Denmark: 1977; Iceland: 1266; Finland: 800; UK: 569; and US: 202), Andersen et al. (2025) identified eight independent genetic variants—six common and two rare—associated with HS. Four of these variants implicate genes in the Notch and Wnt/β-catenin signaling pathways, which regulate epidermal keratinization and hair follicle biology. The study provides compelling evidence that hyperkeratinization, rather than immune dysregulation alone, may be central to HS pathogenesis.
Introduction
HS affects approximately 1% of the global population, with higher prevalence in certain regions. It is characterized by painful nodules, abscesses, and sinus tracts in intertriginous areas. While environmental factors like obesity and smoking are known contributors, the genetic basis of HS has remained elusive. Previous studies identified rare mutations in γ-secretase complex genes, but these account for a small fraction of cases. This study aimed to uncover both common and rare variants contributing to HS risk and to elucidate the biological pathways involved.
Methodology
The meta-GWAS combined data from Denmark, Iceland, Finland, the UK, and the US. Logistic regression was used to test associations, adjusting for confounders and population stratification. Bonferroni correction was applied for genome-wide significance. Variant-to-gene mapping included eQTL and pQTL analyses, structural modeling (AlphaFold), and functional enrichment using STRING and Cytoscape.
Key Findings
- Eight independent variants were associated with HS, including:
- Rare variants in NCSTN (stop-gain) and PSENEN (frameshift), both γ-secretase components.
- Common variants in WNT10A (missense), TMED10 (eQTL), and intergenic regions near SOX9 and KLF5.
- Functional impact:
- WNT10A mutation (p.Phe228Ile) affects Wnt signaling and is linked to thin skin phenotypes.
- TMED10 variant reduces expression of a γ-secretase inhibitor, potentially increasing Notch signaling.
- NCSTN and PSENEN mutations disrupt epidermal development during embryogenesis.
- Pathway enrichment:
- Significant enrichment in Notch and Wnt/β-catenin pathways.
- Associated proteins involved in keratinocyte differentiation, desquamation, and skin barrier function.
- Replication:
- Two previously reported variants near SOX9 and KLF5 were confirmed.
Critical Analysis
Strengths
- Large sample size and multi-national cohorts enhance statistical power and generalizability (within Caucasian populations).
- High-resolution sequencing enabled detection of rare variants.
- Comprehensive functional annotation strengthens biological plausibility.
Limitations
- Limited racial diversity restricts applicability to non-Caucasian populations.
- Lack of clinical severity data (e.g., Hurley staging) prevents genotype-phenotype correlation.
- No direct functional validation of variant effects in vitro or in vivo.
Implications
This study shifts the paradigm of HS pathogenesis from immune dysregulation to epidermal hyperkeratinizationdriven by genetic variants in key signaling pathways. It suggests that keratinolytic therapies may be more effective than immunosuppressants in some patients. Moreover, the findings open avenues for biomarker development and precision medicine approaches targeting Notch and Wnt signaling.
Conclusion
The identification of both common and rare variants in genes regulating keratinization provides a genetic framework for understanding HS. The involvement of Notch and Wnt/β-catenin pathways underscores the importance of epithelial biology in disease onset and progression. Future studies should explore these mechanisms in diverse populations and assess therapeutic interventions targeting these pathways.
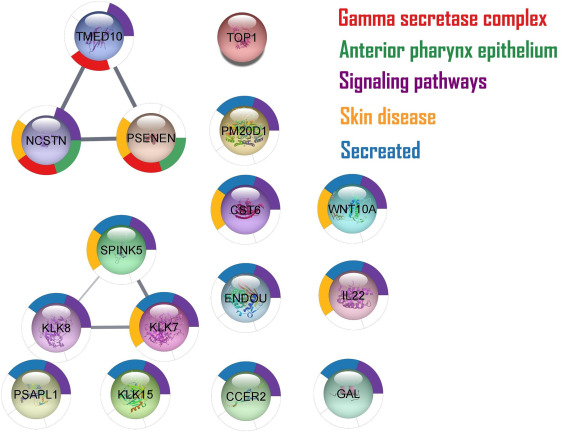
Fig 2 Functional enrichment analyses for the gene products associated with the HS phenotype. Functional enrichment analysis performed on candidate causative genes at the locus, in combination with protein quantitative trait loci affected at the locus as determined after variant to gene mapping. The enrichment was performed via STRING v11 and the Cytoscape 3.9.1 software in combination with the functional enrichment tool in the stringApp. Depictured are the 11 gene products found to be associated with the HS phenotype and color coded for the cellular components, cellular pathways, tissue localization, or known diseases enriched for within these gene products. The multicolored circle surrounding the individual products represents the areas for which the network achieved functional enrichment within the areas of cellular components, cellular pathways, tissue localization, and known diseases. The interconnecting lines indicate association between the genes in medical literature, with the bold lines indicating direct association as validated through pull-down analyses (in this figure representing, eg, the γ-secretase complex). It should be noted that while KLK8 did not appear as part of the enrichment for “skin diseases” both KLK8 and SPINK5 were statistically significantly enriched for “Netherton syndrome” (FDR = 0.0119)—a rare but well-known skin disease. CCER2, Coiled-coil glutamate rich protein 2; CST6, cystatin M; ENDOU, uridylate-specific endoribonuclease; GAL, galanin; HS, hidradenitis suppurativa; IL22, interleukin-22; KLK7, serine protease Kallikrein-7; KLK8, serine protease Kallikrein-8; KLK15, serine protease Kallikrein-15; NCSTN, nicastrin; PM20D1, peptidase M20 domain-containing protein 1; PSENEN, presenilin enhancer protein 2; PSAPL1, proactivator polypeptide-like 1; SPINK5, serine protease inhibitor Kazal-type 5; TMED10, transmembrane P24 trafficking protein 10; TOP1, DNA topoisomerase I; WNT10A, wingless/integrated 10A.
Sources
- Andersen RK et al. A genome-wide association meta-analysis links hidradenitis suppurativa to common and rare sequence variants causing disruption of the Notch and Wnt/β-catenin signaling pathways. JAAD. 2025;92(4):761–772. DOI: 10.1016/j.jaad.2024.11.050
- Supplementary data: Mendeley Dataset
- GTEx Portal: https://www.gtexportal.org/home/
- AlphaFold Protein Structure Database: https://alphafold.ebi.ac.uk/
- STRING Database: https://string-db.org/